Code&Data
We strive to make all of our tools, protocols and datasets available to the community. For the most up to date source codes, visit our group page on Github.
Source code
We are building computatational pipelines for weakly supervised image analysis in digital pathology. We strive to always incorporate the latest technologies, so our pipelines are constantly evolving.
- Our current standard pipeline for weakly supervised pathology image analysis is “marugoto” (2022-2023): https://github.com/KatherLab/marugoto
- “Deepmed” package by Marko van Treeck (Python implementation, 2020-2022): https://github.com/KatherLab/deepmed
- “Histology image analysis” (HIA) package by Narmin Ghaffari Laleh (Python implementation, 2020-2022): https://github.com/KatherLab/HIA
- Deep learning-based prediction of molecular alterations pan-cancer package (Matlab implementation, 2019-2021): https://github.com/jnkather/DeepHistology
- Deep learning for detecting virus presence in cancer images, from 2018/2019: https://github.com/jnkather/VirusFromHE
- Deep learning for detecting MSI in gastrointestinal cancer, original codes from 2018/2019: https://github.com/jnkather/MSIfromHE
Metadata
- Metadata for the TCGA cohort, preprocessed for computational pathology analyses: https://github.com/KatherLab/cancer-metadata
Trained models
- our latest models for MSI prediction in colorectal cancer (PyTorch) are available at https://zenodo.org/record/5151502
Cancer histology images
Human solid tumors are made up of many different tissue types. Image analysis pipelines often start with a classification of these regions (such as tumor, stroma, necrosis, etc.). These are labeled, quality-controlled sets of images that can be used to train tissue classifiers:
- Benchmark data sets - we show the functionalities of Deepmed on two benchmark datasets, TCGA-BRCA-A2 and TCGA-BRCA-E2, that are available at https://zenodo.org/record/5337009
- 5000 labeled images of colorectal cancer tissue (from this paper): download
- 100,000 labeled images of colorectal cancer tissue (from this paper): download
- 1,000,000 images of colorectal cancer tissue in: download
- ˜12k images for tumor detection in colorectal and gastric cancer (512x512 px at 0.5 µm/px, from this paper): download
After detecting tissue of interest in whole slide images, deep learning classifiers can extract clinically meaningful information from the images. These datasets can be used to train these classifiers:
-
˜400k image patches of microsatellite instable (MSI) vs. microsatellite stable (MSS) image patches of colorectal and gastric cancer (from this paper): download, derived from the TCGA data set at http://portal.gdc.cancer.gov.
-
image patches of all colorectal cancer (CRC) whole slide images from the TCGA database, conveniently cut into tiles of 512x512 px for subsequent deep learning analysis. Only the manually annotated tumor region was processed. Patient pseudonyms (TCGA barcodes) are preserved in the dataset: https://zenodo.org/record/3784345. Corresponding genetic information are available at https://cbioportal.org. Original data credit: http://portal.gdc.cancer.gov.
Generated images
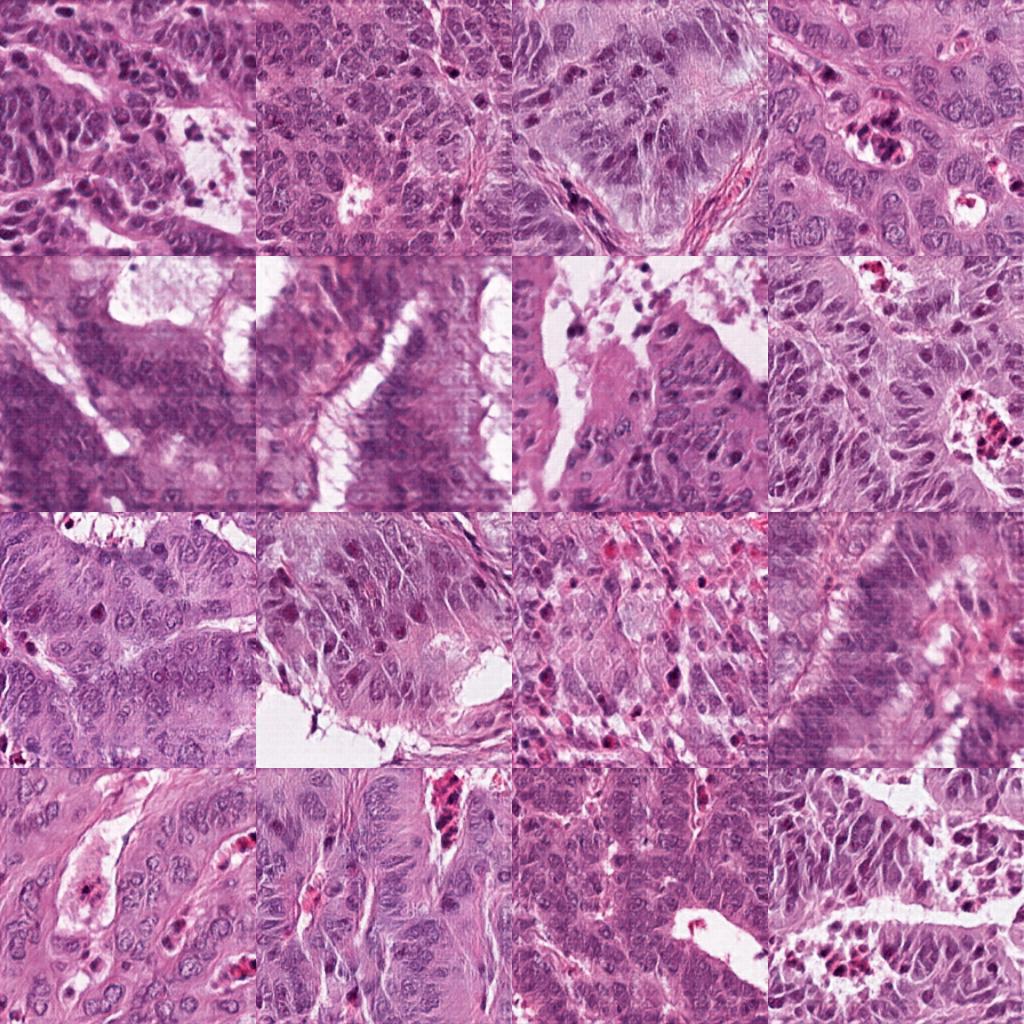
Deep generative adversarial networks can generate realistic histology images:
- 2500 image tiles of generated colorectal cancer tissue, 256x256 px
Protocols
- The “Aachen Protocol” for data preprocessing in deep learning histology image analysis: https://zenodo.org/record/3694994
Others
- Manual tumor annotations on TCGA diagnostic slides: https://zenodo.org/record/5320076
- trained Pytorch models for MSI/dMMR status prediction in colorectal cancer: https://zenodo.org/record/5151502